This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison
Microarray
Microarrays have become crucial in recent years to look at differing gene expression in cells not only between disease and non-disease states but also at differing gene expression throughout the day. This is done by first placing onto a chip thousands of differing cDNA sequences, which represent the genome mRNAs are then tagged either green from a diseased organism or red from a non-diseased organism, these mRNA florescence tags can be either way. The two mRNA batches are then washed over separate identical chips, which are then photographed and visually overlaid on top of one another. Gene expression can then be quantified as either turned on either in the disease state, gene highlighted as green, non-disease state, red, or in both states the gene might have similar expression, seen as black on the microarray. As I stated earlier it is also possible to do a microarray analysis of genes at different points of the day, this screen would follow the same parameters, however the two mRNAs would come from a day and a night sampled organism not disease and non-diseased state organisms.
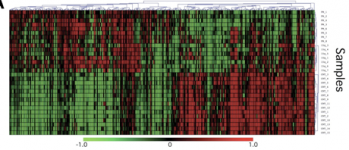
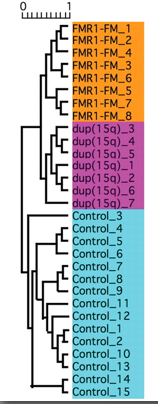
Fragile X syndrome, as stated previously, is a lack of the FMR1 protein, therefore looking initially at gene expression there would be a change in the FMR1 gene’s expression between a FXS patient and a patient with a functioning FMR1 gene. However as curious scientists we also want to know what other genes have a change in expression in the diseased state. Below you can see one representative of a microarray comparing multiple FMR1 mutants, duplications of chromosome 15-another from of autism, and non-autistic humans.
Fragile X syndrome, as stated previously, is a lack of the FMR1 protein, therefore looking initially at gene expression there would be a change in the FMR1 gene’s expression between a FXS patient and a patient with a functioning FMR1 gene. However as curious scientists we also want to know what other genes have a change in expression in the diseased state. Below you can see one representative of a microarray comparing multiple FMR1 mutants, duplications of chromosome 15-another from of autism, and non-autistic humans.
It can be seen from the microarray above that if the sample was either a FMR1 mutation or a duplication of 15q11-q13 gene expression was opposite to the control genes without autism. The enhancement of expression when the controls were repressed might show that those genes might have been needed to compensate for the missing genes. It should also be noted that while for a majority of genes the FMR1 FXS patients and the 15q11-13 duplications has almost identical gene expression there were some genes that did differ between the two, as autism is caused by many different genes other than FMR1, and FXS patients are not solely autistic.
In conclusion, while microarrays help show gene expression in an organism, it should be noted with complex diseases a microarray can often open more questions than it will solve. They can be great indicators at what genes can be ignored or should be given more emphasis. However even with their vast power and ease they should be used in addition to other methods to strengthen their power and answer questions they ignite.
In conclusion, while microarrays help show gene expression in an organism, it should be noted with complex diseases a microarray can often open more questions than it will solve. They can be great indicators at what genes can be ignored or should be given more emphasis. However even with their vast power and ease they should be used in addition to other methods to strengthen their power and answer questions they ignite.
References
[1] GEO: www.ncbi.nlm.nih.gov/geo
[2] http://www.ncbi.nlm.nih.gov/geo/gds/profileGraph.cgi?&dataset=njrzO3gliogZ0ek-AJiffmmOT5_kgP&dataset=rmtvR8noprmZ1kohIKqmnrrOSb_rmO$&gmin=-0.220000&gmax=0.263000&absc=&gds=2824&idref=23816&annot=A_24_P127661
[3] Yuhei Nishimura, Christa L. Martin, et. al. Genome-wie expression profiling of lymphoblastoid cell lines distinguishes differenct forms of autism nd reveals shared pathways. Human Molecular Genetics Advance Access. July 15, 2007. Hum. Mol. Genet. 16: 1682-1698
[2] http://www.ncbi.nlm.nih.gov/geo/gds/profileGraph.cgi?&dataset=njrzO3gliogZ0ek-AJiffmmOT5_kgP&dataset=rmtvR8noprmZ1kohIKqmnrrOSb_rmO$&gmin=-0.220000&gmax=0.263000&absc=&gds=2824&idref=23816&annot=A_24_P127661
[3] Yuhei Nishimura, Christa L. Martin, et. al. Genome-wie expression profiling of lymphoblastoid cell lines distinguishes differenct forms of autism nd reveals shared pathways. Human Molecular Genetics Advance Access. July 15, 2007. Hum. Mol. Genet. 16: 1682-1698
Aime Agather
[email protected]
Last Updated 05/16/2010
Genetics 677
[email protected]
Last Updated 05/16/2010
Genetics 677