This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison
Motifs from STAMP

Familial Profile for GATA Motifs from STAMP
Initially human FMR1 domains were isolated with help of MotifFinder [1] and scanned for repeated domains. Initially at a cutoff value of 85, 62 motifis were isolated from the FMR1 DNA sequence. When I increased to the cutoff value to 95 out of the 7 sequences, 2 of which were heat shock factors. While heat shock factors do have biological significance, I decided that I would not analyze these domains, as protein integrity at high temperatures is a common theme for all proteins across all organisms.
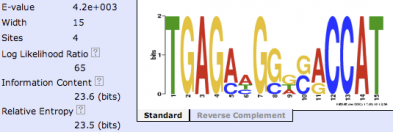
At a cut off value of 90 there were 25 different motifs, of which I decided to look at repeating GATA binding domains, as these are crucial for binding proteins to DNA or mRNA [2], a main component biological process for the FMR1 protein [3]. Check out the gene ontology page for more information on the cellular components, molecular functions and biological processes of the FMR1 protein. Looking at the familial motif sequence fidelity you can see the high levels of conservation by the size of the each letter in the sequence. Three of the six main letters of the motifs were solely A, T, and C. This high fidelity of the GATA DNA sequences along the protein, transport of mRNA is important for some function in neuronal cells.
At a cut off value of 90 there were 25 different motifs, of which I decided to look at repeating GATA binding domains, as these are crucial for binding proteins to DNA or mRNA [2], a main component biological process for the FMR1 protein [3]. Check out the gene ontology page for more information on the cellular components, molecular functions and biological processes of the FMR1 protein. Looking at the familial motif sequence fidelity you can see the high levels of conservation by the size of the each letter in the sequence. Three of the six main letters of the motifs were solely A, T, and C. This high fidelity of the GATA DNA sequences along the protein, transport of mRNA is important for some function in neuronal cells.
Famillial profile for the GATA domains
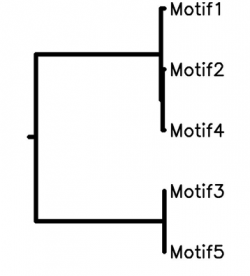
To the right is the result from STAMP, a program that compares motif similarities and makes a phylogeny map.
Motifs 1 and 3 are GATA-binding factor 1s
Motif 2 is GATA-binding factor 2
Motif 4 is a GATA-binding factor 3
Motif 5 is a GATA binding site
Given the relations between the motifs I find it interesting that the two GATA-binding factor 1s are not as closely related to one another but rather one is closer in its evolutionary history to the binding site while the other is related to the two other binding factors.
Motifs 1 and 3 are GATA-binding factor 1s
Motif 2 is GATA-binding factor 2
Motif 4 is a GATA-binding factor 3
Motif 5 is a GATA binding site
Given the relations between the motifs I find it interesting that the two GATA-binding factor 1s are not as closely related to one another but rather one is closer in its evolutionary history to the binding site while the other is related to the two other binding factors.
Motifs from MEME
MEME [5] was another program that was available to help identify motifs in DNA sequences. Once I put the FMR1 DNA sequence MEME only gave three motifs out. Below are the three motifs identified from the FMR1 gene and their locations in the gene.
STAMP vs. MEME
The main difference between MEME and MotifFinder is that MotifFinder characterized and named all of the motifs the program identified in the FMR1 DNA sequence. In my opinion I thought that was helpful so anyone using it can identify a motif they want to look at further. The one thing I thought MEME excelled at over MotifFinder was that the results gave the multiple alignments of the DNA sequence between the different motifs right away so visual learners, like myself, could see what motifs had the highest fidelity along the gene. However while using MEME you had to then identify what each motif meant by searching through another program, MAST, the website lists. This seems rather superfluous and the motifs' identities could have been included on the output from MEME.
References
[1] MotifFinder
[2] Cheng, Y., King, D. C., Dore, L. C., Zhang, X., Zhou, Y., Zhang, Y. et al. (2008). Transcriptional enhancement by GATA1-occupied DNA segments is strongly associated with evolutionary constraint on the binding site motif. Genome Research, 18(12), 1896-1905.
[3] AmiGO
[4] STAMP http://www.benoslab.pitt.edu/stamp/
[5] MEME
[2] Cheng, Y., King, D. C., Dore, L. C., Zhang, X., Zhou, Y., Zhang, Y. et al. (2008). Transcriptional enhancement by GATA1-occupied DNA segments is strongly associated with evolutionary constraint on the binding site motif. Genome Research, 18(12), 1896-1905.
[3] AmiGO
[4] STAMP http://www.benoslab.pitt.edu/stamp/
[5] MEME
Aime Agather
[email protected]
Last Updated 03/28/2010
Genetics 677
[email protected]
Last Updated 03/28/2010
Genetics 677




